 |
|
|
|
| Model Performance | ||||||||||||||||||||||||
|
|
||||||||||||||||||||||||
|
||||||||||||||||||||||||
| Nearest Neighbors from Test Set |
|
|
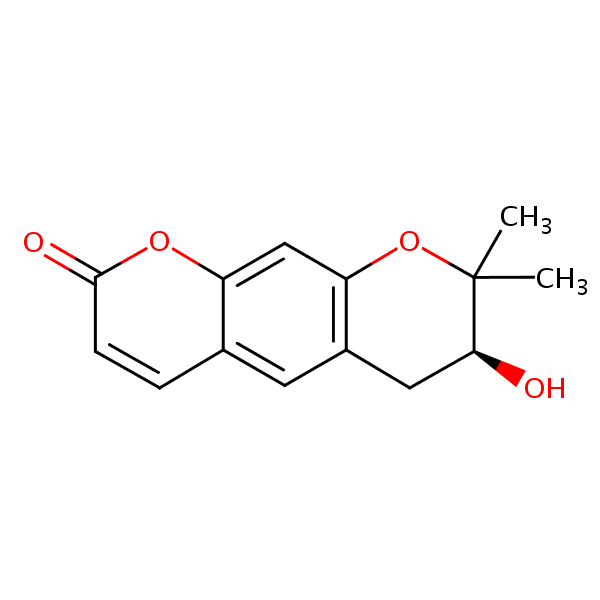
| Test chemical Measured: N/A Predicted: -7.12  Decursinol |
Similarity: 0.68 Measured: -12.0 Predicted: -12.1 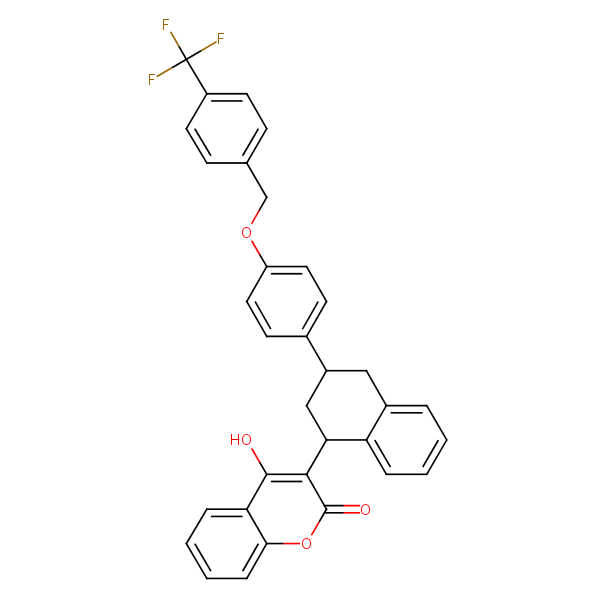 Flocoumafen |
Similarity: 0.56 Measured: -7.59 Predicted: -7.26  Fenazaquin |
Similarity: 0.54 Measured: -4.85 Predicted: -6.70  Chlomethoxyfen |
Similarity: 0.53 Measured: -7.23 Predicted: -9.11 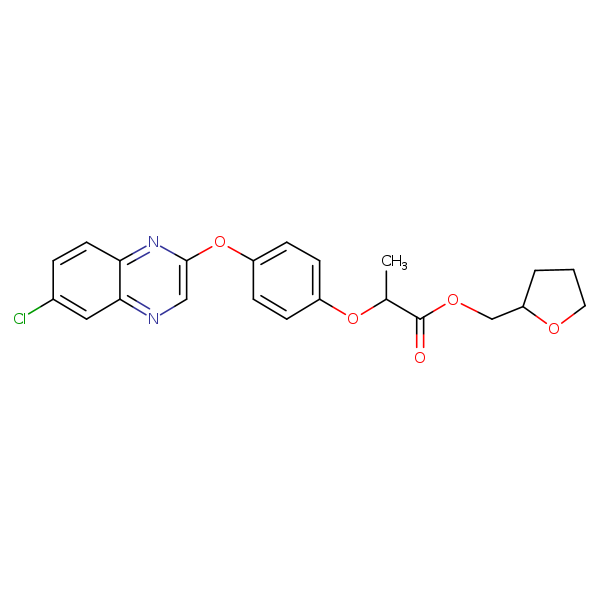 Quizalofop-tefuryl |
Similarity: 0.53 Measured: -8.13 Predicted: -8.48 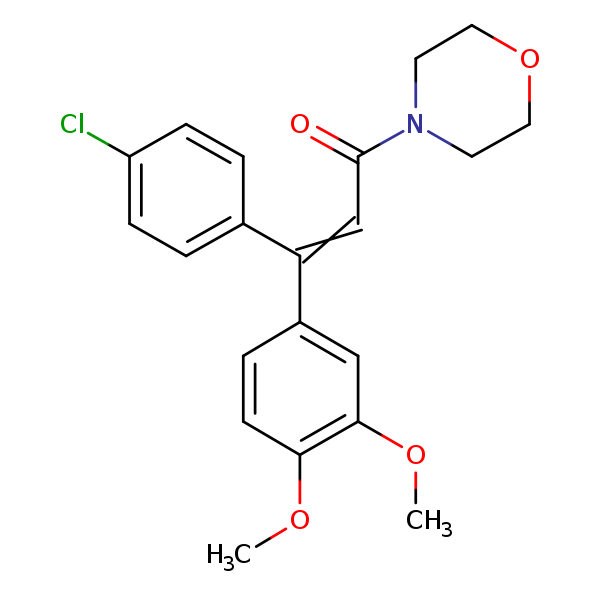 Dimethomorph |
| Similarity: 0.52 Measured: -9.06 Predicted: -9.32 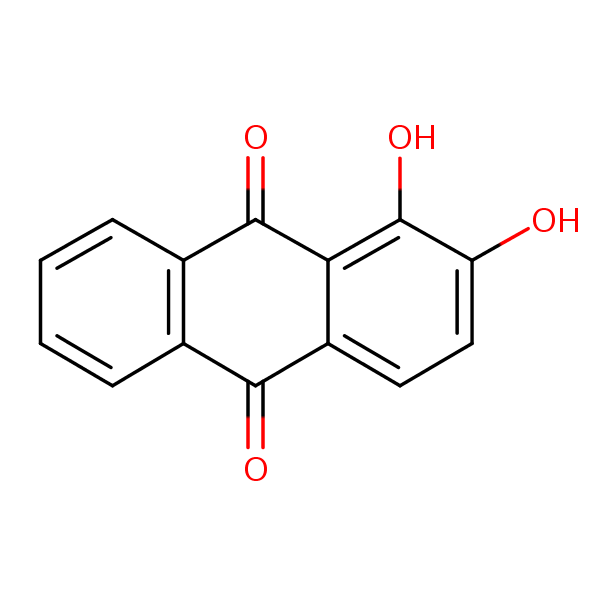 Alizarin |
Similarity: 0.51 Measured: -5.00 Predicted: -6.47  Benalaxyl |
| Nearest Neighbors from Training Set |
|
|
| Test chemical Measured: N/A Predicted: -7.12  Decursinol |
Similarity: 0.69 Measured: -3.01 Predicted: -2.31 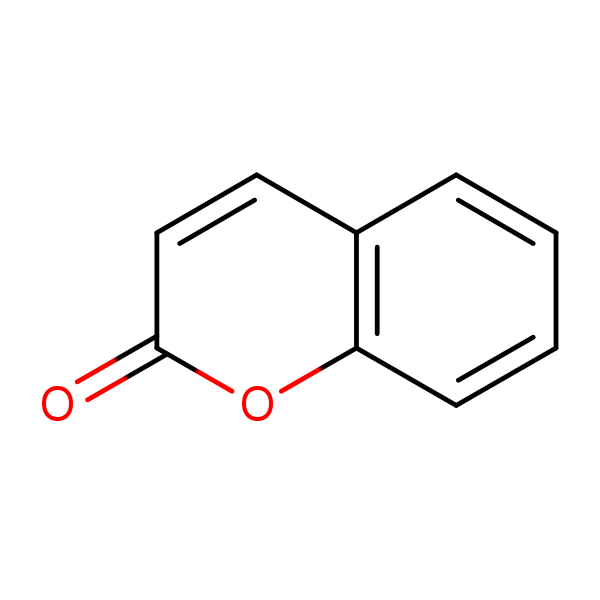 Coumarin |
Similarity: 0.57 Measured: -7.40 Predicted: -6.85 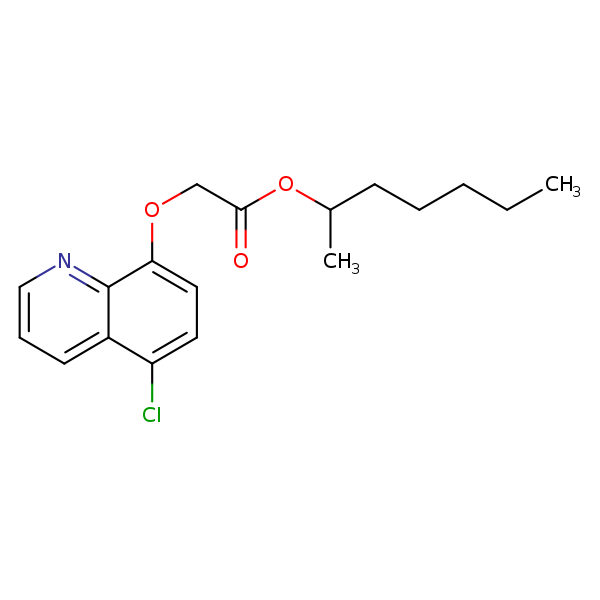 Cloquintocet-mexyl |
Similarity: 0.56 Measured: -10.2 Predicted: -7.88 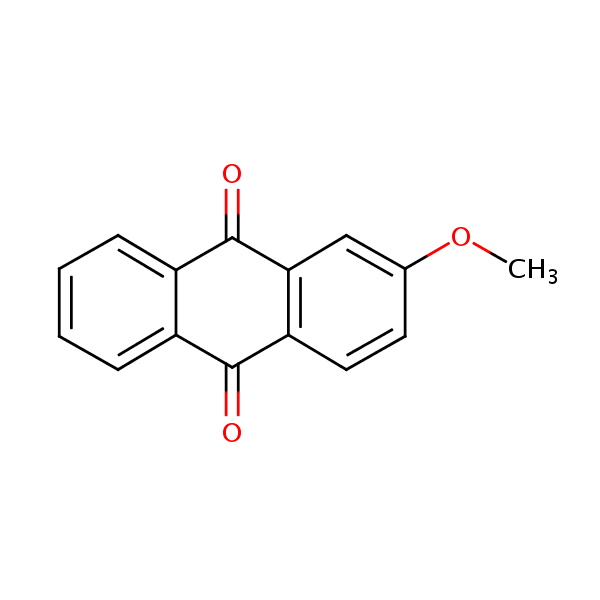 9,10-Anthracenedione, 2-methoxy- |
Similarity: 0.55 Measured: -11.0 Predicted: -9.72 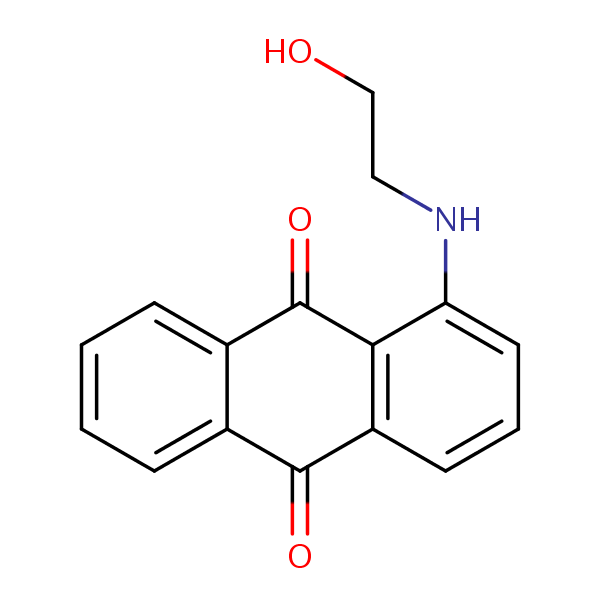 9,10-Anthracenedione, 1-[(2-hydroxyethyl... |
Similarity: 0.55 Measured: -5.46 Predicted: -6.25 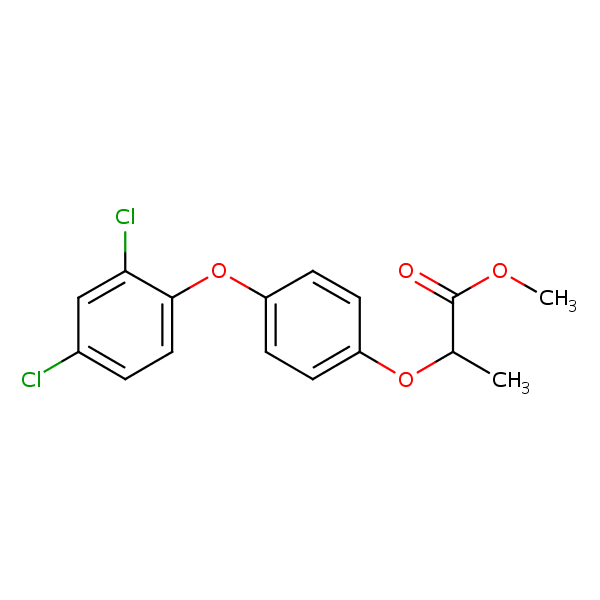 (+-)-Diclofop-methyl |
| Similarity: 0.55 Measured: -6.82 Predicted: -6.50 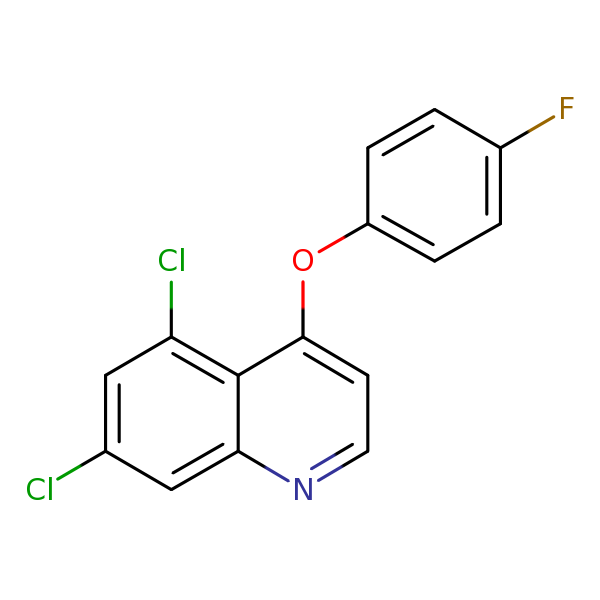 Quinoxyfen |
Similarity: 0.54 Measured: -6.56 Predicted: -7.69 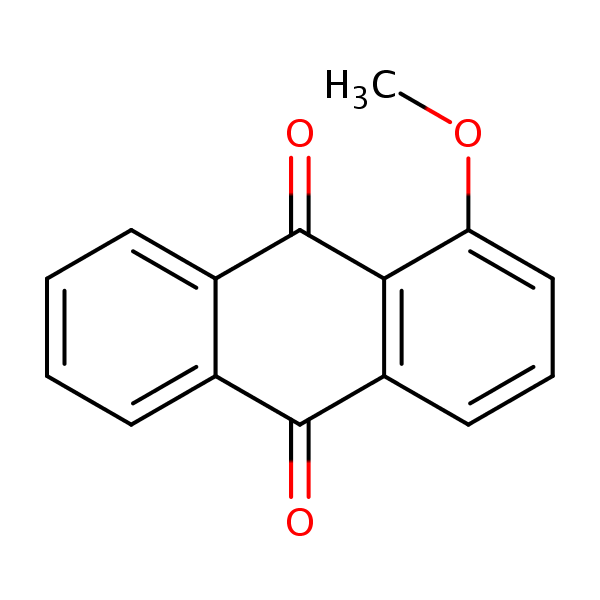 9,10-Anthracenedione, 1-methoxy- |
Similarity: 0.53 Measured: -6.76 Predicted: -6.94 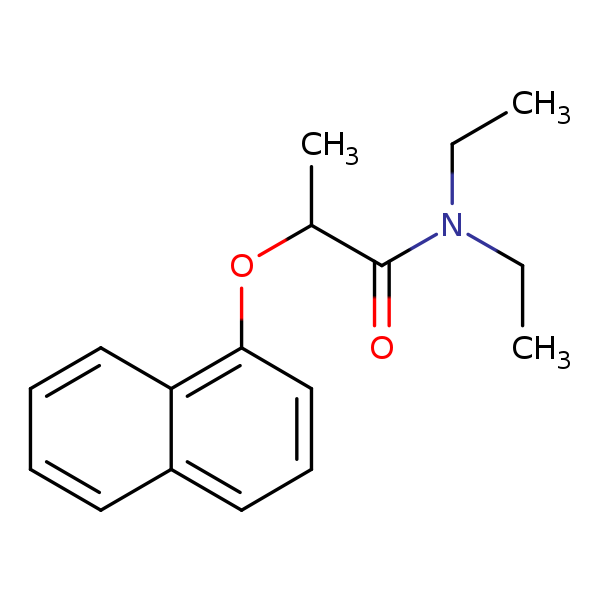 Napropamide |
Similarity: 0.53 Measured: -8.23 Predicted: -7.52 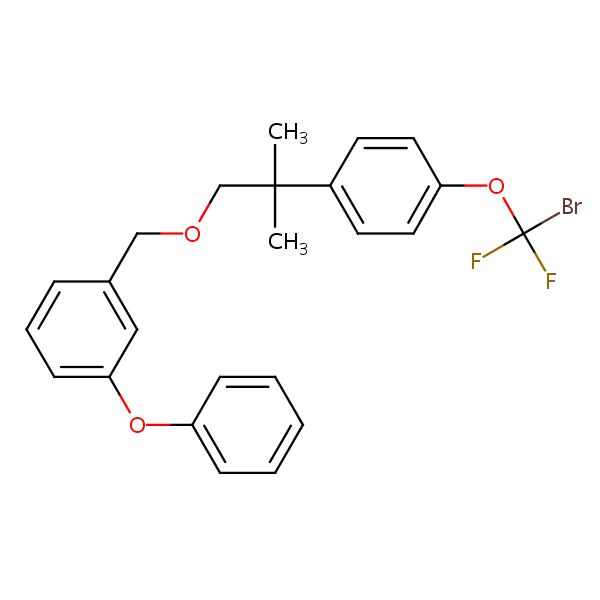 Halfenprox |
Similarity: 0.53 Measured: -10.2 Predicted: -8.70 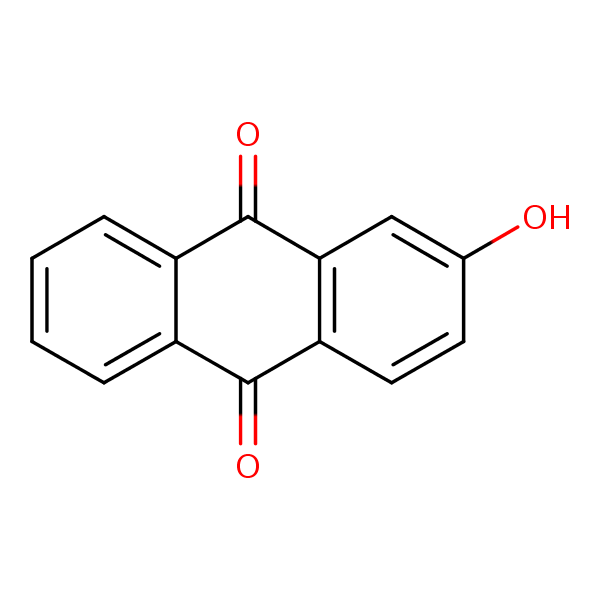 2-Hydroxyanthraquinone |