 |
|
|
|
| Model Performance | ||||||||||||||||||||||||
|
|
||||||||||||||||||||||||
|
||||||||||||||||||||||||
| Nearest Neighbors from Test Set |
|
|
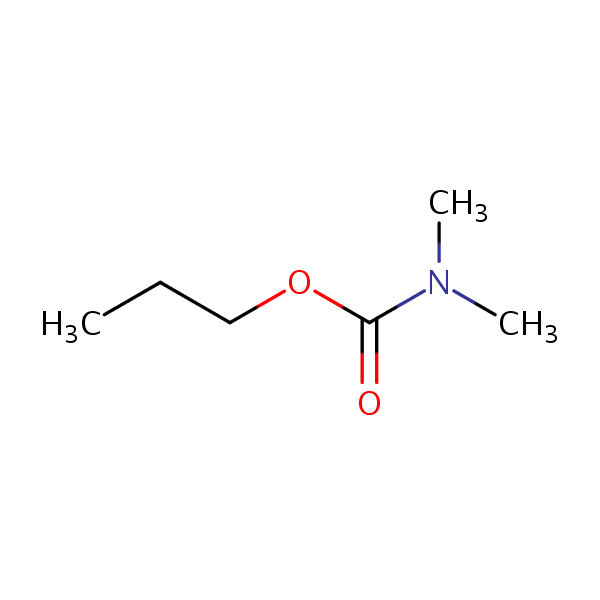
| Test chemical Measured: N/A Predicted: 0.527  Propyl dimethylcarbamate |
Similarity: 0.65 Measured: 0.364 Predicted: 0.623 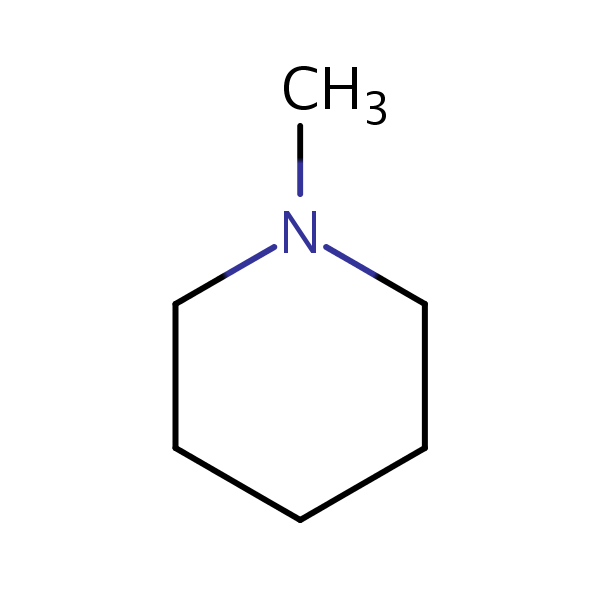 Piperidine, 1-methyl- |
Similarity: 0.65 Measured: 0.321 Predicted: N/A 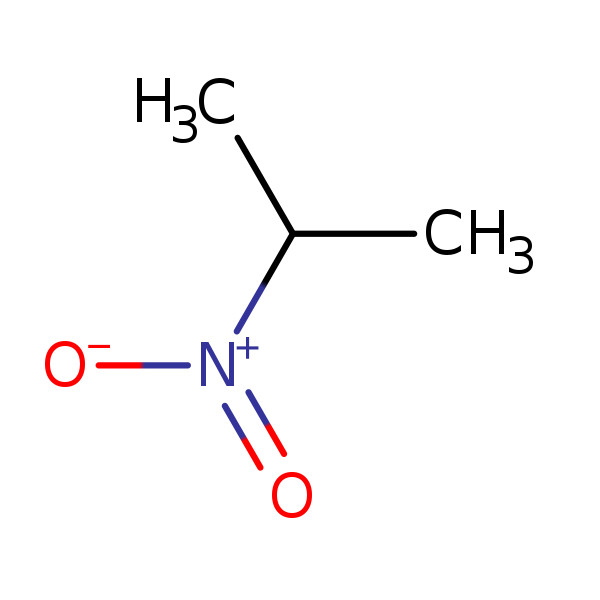 2-Nitropropane |
Similarity: 0.60 Measured: 0.157 Predicted: 0.145 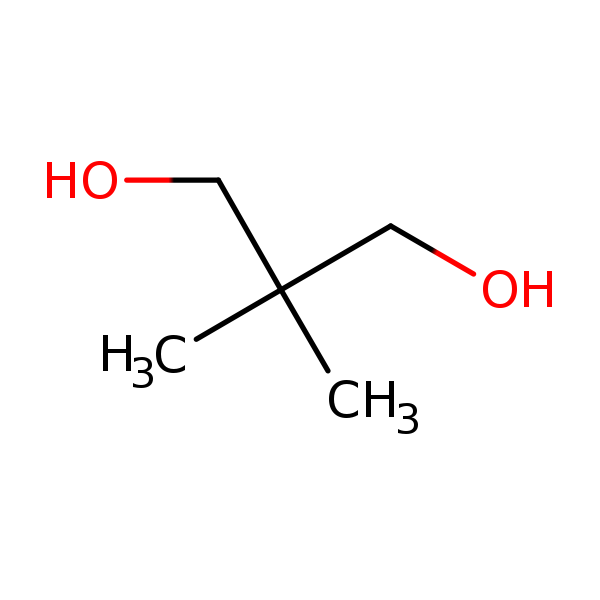 Neopentyl glycol |
Similarity: 0.52 Measured: 1.02 Predicted: 0.375  2-Aminopyridine |
Similarity: 0.52 Measured: 0.301 Predicted: 0.873  1,1,1-Trichloro-2-methyl-2-propanol |
| Similarity: 0.51 Measured: 0.799 Predicted: 0.827  Chloroform |
| Nearest Neighbors from Training Set |
|
|
| Test chemical Measured: N/A Predicted: 0.527  Propyl dimethylcarbamate |
Similarity: 0.71 Measured: -0.107 Predicted: -0.167  N,N-Dimethylformamide |
Similarity: 0.68 Measured: 0.0955 Predicted: 0.168  2,2-Dimethylpropanoic acid |
Similarity: 0.64 Measured: 1.91 Predicted: 1.22  Butyl ether |
Similarity: 0.61 Measured: 0.195 Predicted: 0.650  Triethylamine |
Similarity: 0.61 Measured: 1.21 Predicted: 1.09  1-Chlorobutane |
| Similarity: 0.60 Measured: 0.948 Predicted: 1.14  N,N-Dimethylbenzylamine |
Similarity: 0.59 Measured: 0.180 Predicted: 0.613 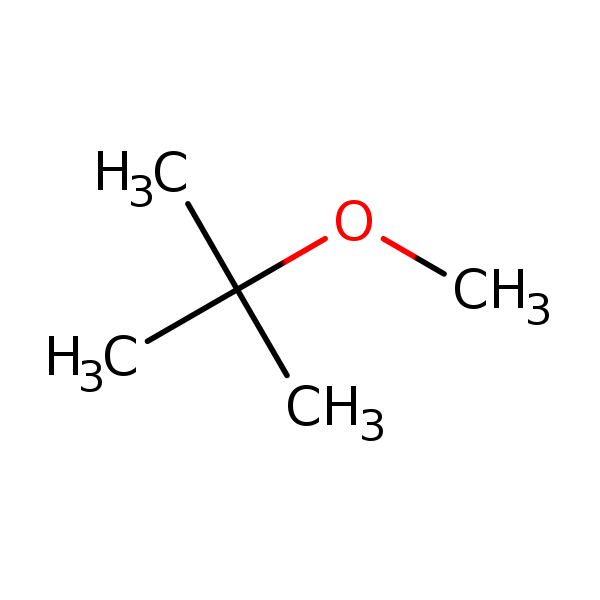 Methyl tert-butyl ether |
Similarity: 0.59 Measured: 0.475 Predicted: 0.502 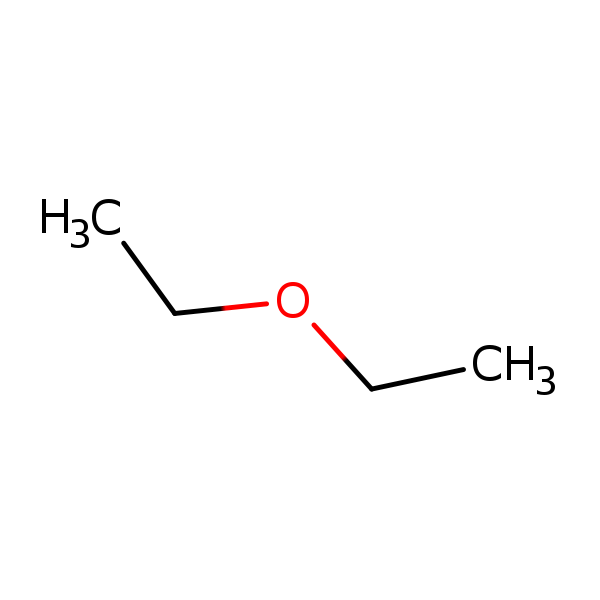 Diethyl ether |
Similarity: 0.58 Measured: 0.295 Predicted: 0.908 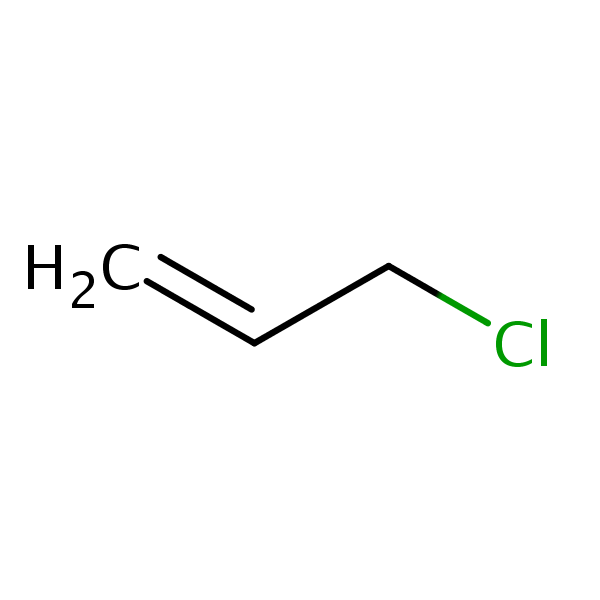 Allyl chloride |
Similarity: 0.58 Measured: 0.664 Predicted: 0.113  Dalapon |