 |
|
|
|
| Model Performance | ||||||||||||||||||||||||
|
|
||||||||||||||||||||||||
|
||||||||||||||||||||||||
| Nearest Neighbors from Test Set |
|
|
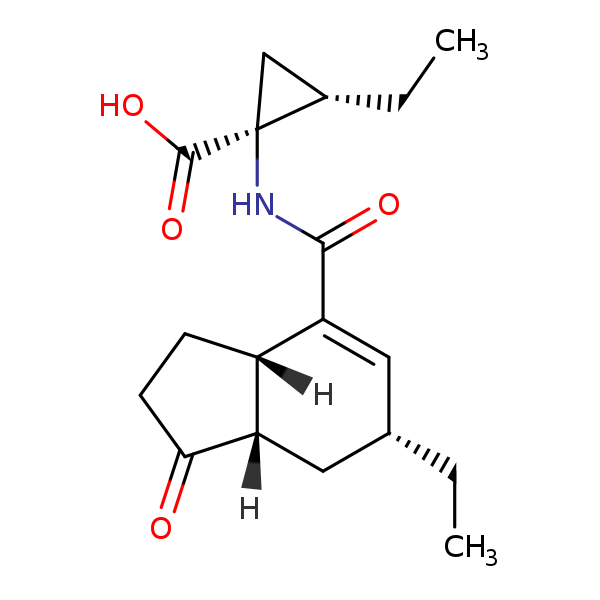
| Test chemical Measured: N/A Predicted: -8.84  Coronatine |
Similarity: 0.78 Measured: -4.69 Predicted: -5.89 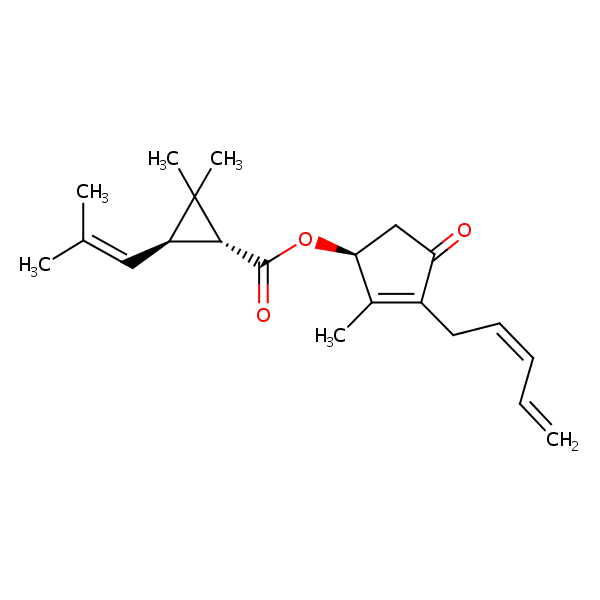 Pyrethrin I |
Similarity: 0.77 Measured: -6.75 Predicted: -6.92 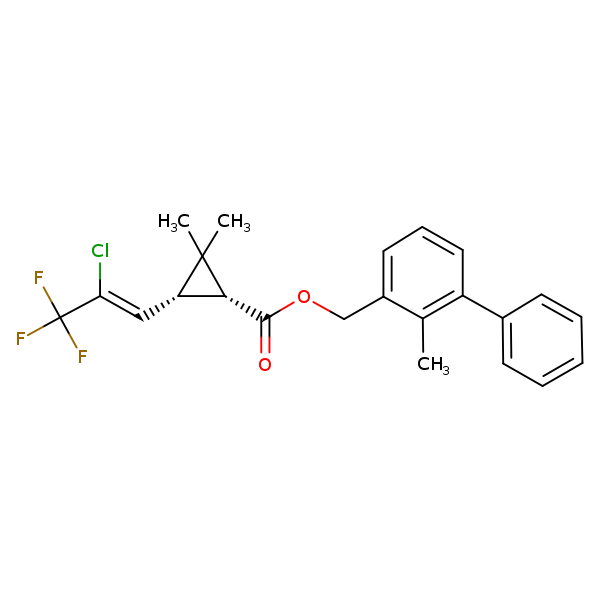 Bifenthrin |
Similarity: 0.73 Measured: -3.85 Predicted: -7.02 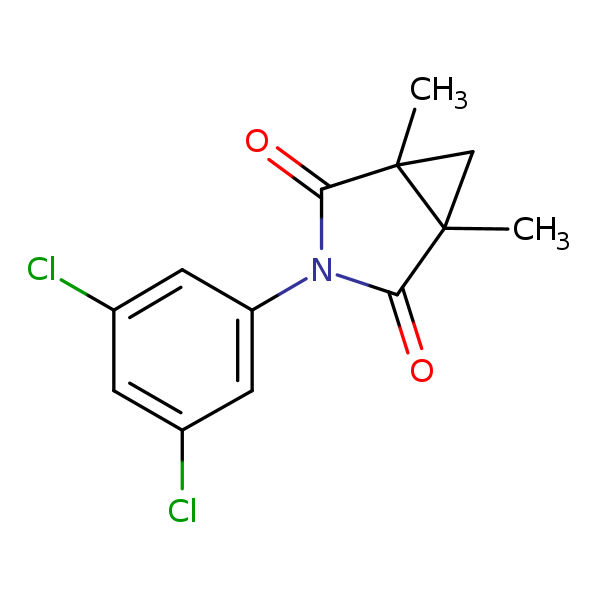 Procymidone |
Similarity: 0.72 Measured: -4.75 Predicted: -7.51 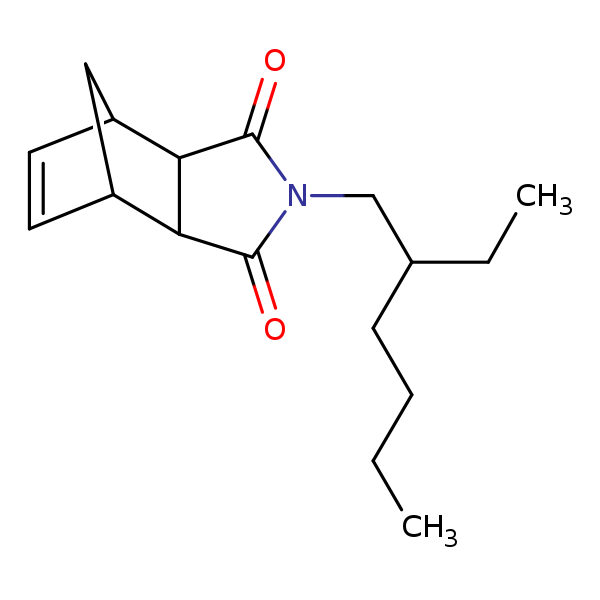 Octylbicycloheptenedicarboximide |
Similarity: 0.63 Measured: -9.13 Predicted: -8.01 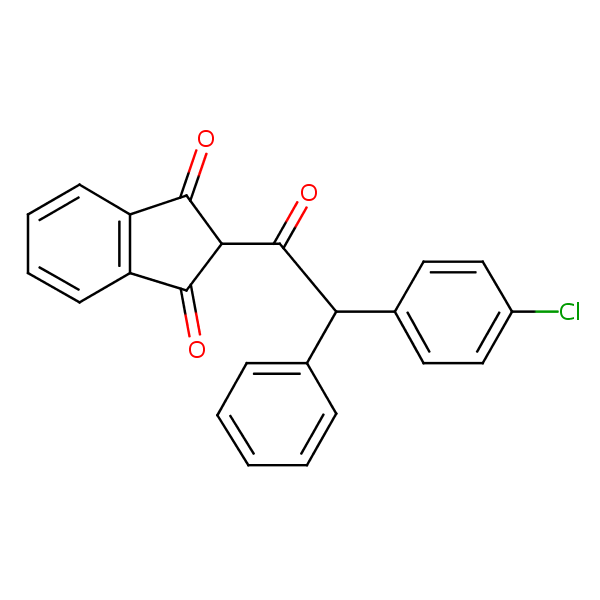 Chlorophacinone |
| Similarity: 0.62 Measured: -9.99 Predicted: -7.38 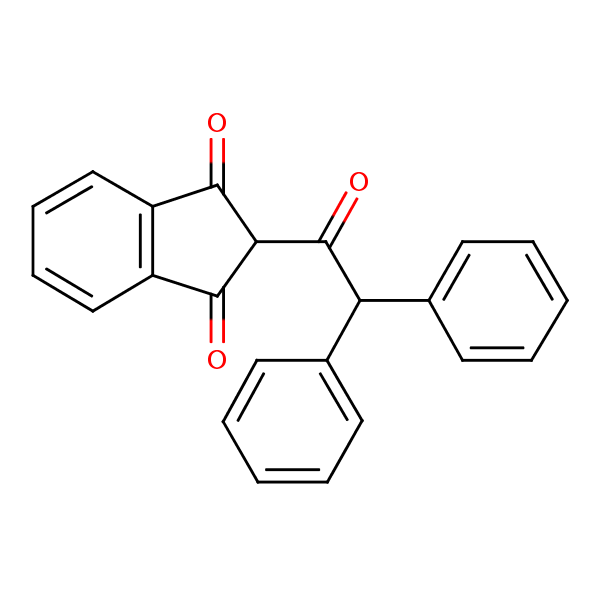 Diphacinone |
Similarity: 0.60 Measured: -6.52 Predicted: -10.1 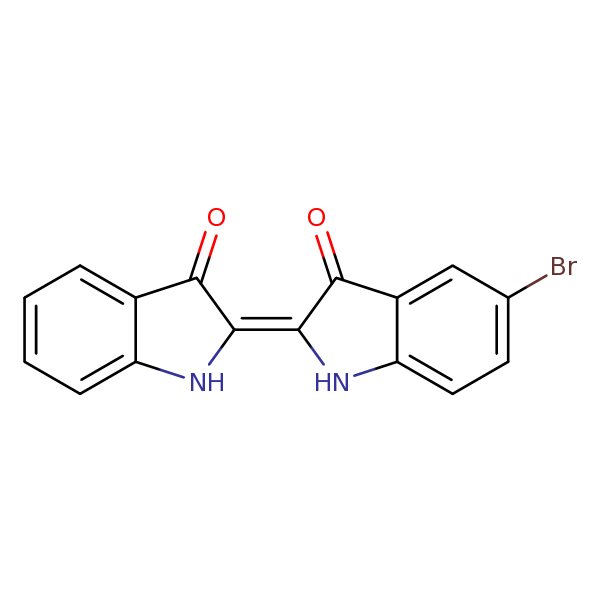 Bromindigo R |
Similarity: 0.60 Measured: -0.385 Predicted: -0.187 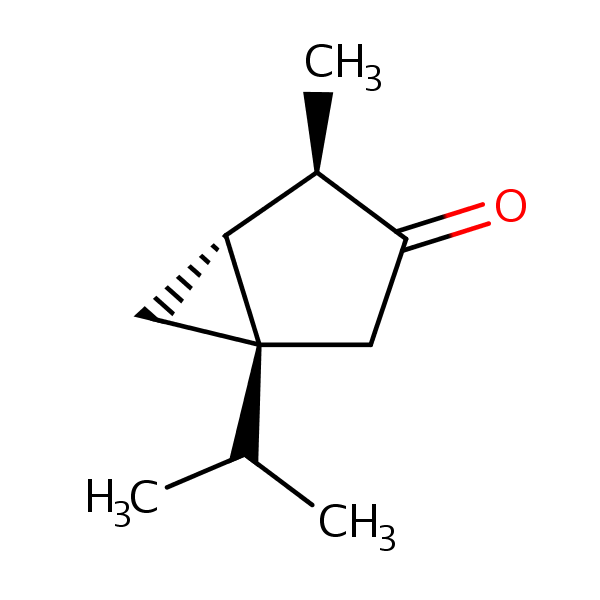 alpha-Thujone |
Similarity: 0.60 Measured: -5.23 Predicted: -5.65 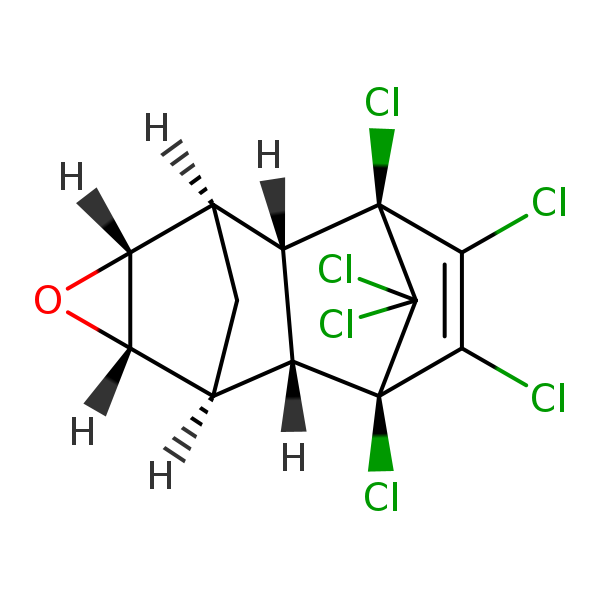 Dieldrin |
Similarity: 0.59 Measured: -5.00 Predicted: -6.47  Benalaxyl |
| Nearest Neighbors from Training Set |
|
|
| Test chemical Measured: N/A Predicted: -8.84  Coronatine |
Similarity: 0.78 Measured: -7.66 Predicted: -8.00 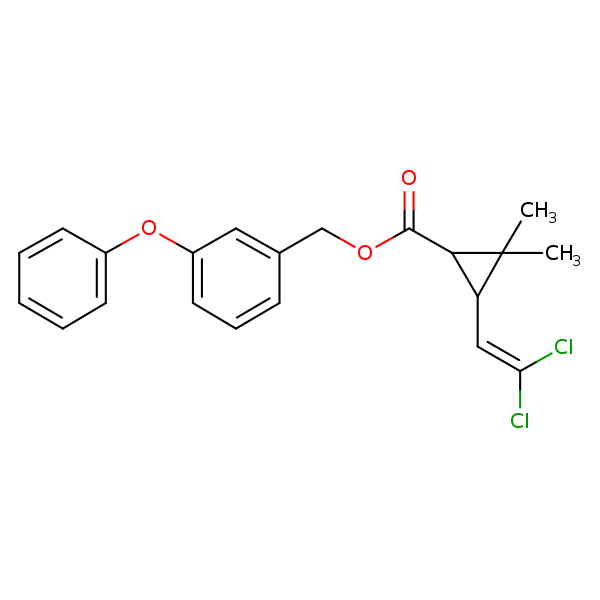 Permethrin |
Similarity: 0.78 Measured: -6.40 Predicted: -6.89 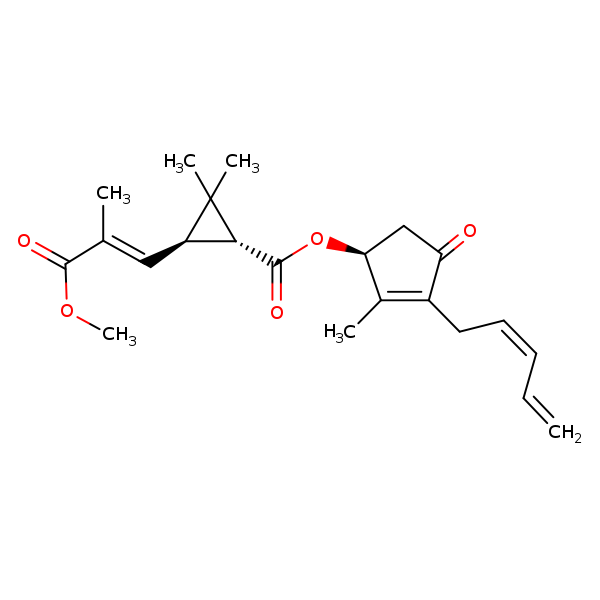 Pyrethrin II |
Similarity: 0.73 Measured: -8.82 Predicted: -8.69 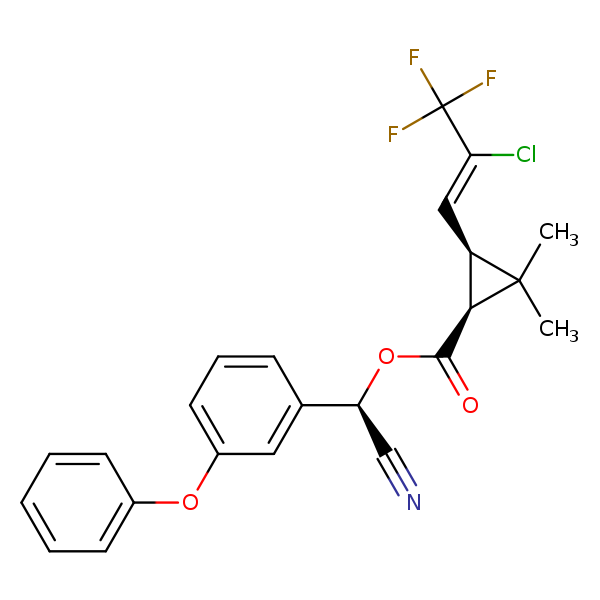 lambda-Cyhalothrin |
Similarity: 0.69 Measured: -7.70 Predicted: -7.82 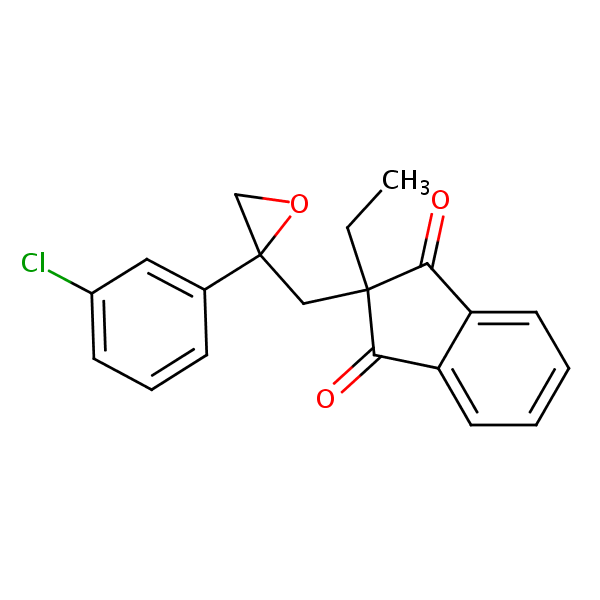 Indanofan |
Similarity: 0.69 Measured: -7.87 Predicted: -7.99 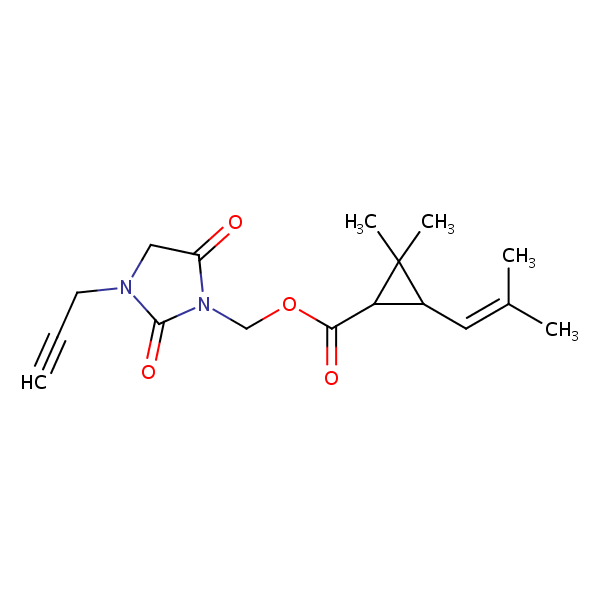 Imiprothrin |
| Similarity: 0.67 Measured: -9.39 Predicted: -9.79 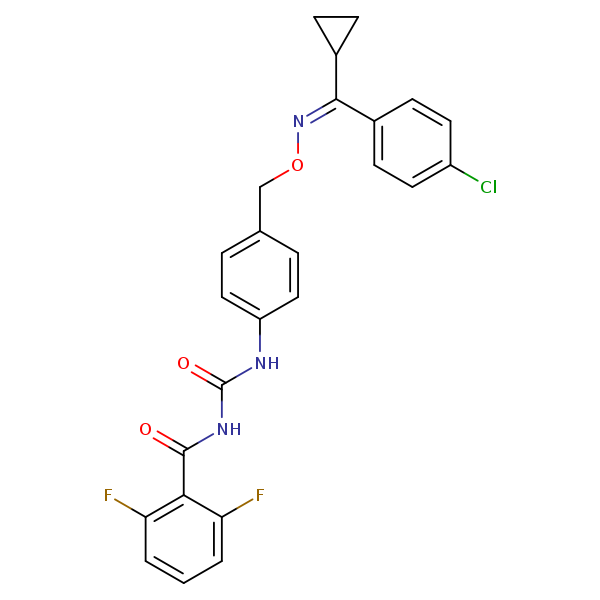 (Z)-Flucycloxuron |
Similarity: 0.66 Measured: -6.70 Predicted: -6.56 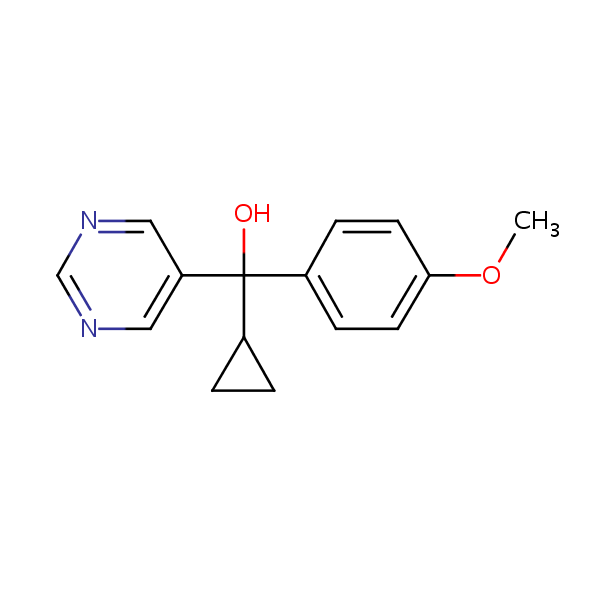 Ancymidol |
Similarity: 0.64 Measured: -10.4 Predicted: -10.1 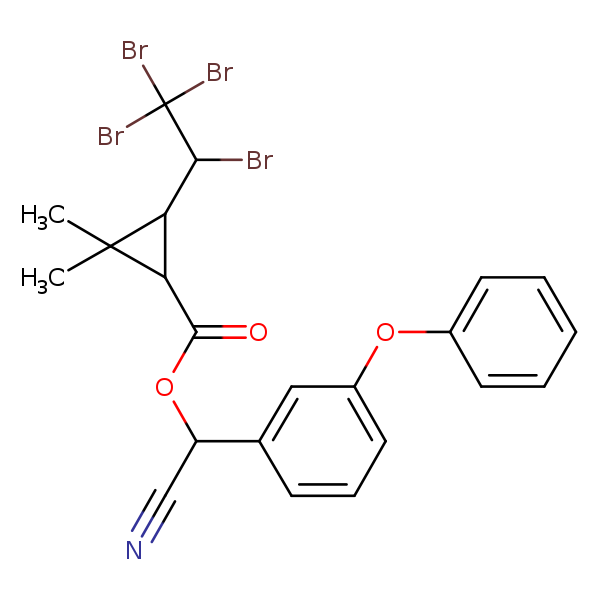 Tralomethrin |
Similarity: 0.64 Measured: -8.82 Predicted: -8.37 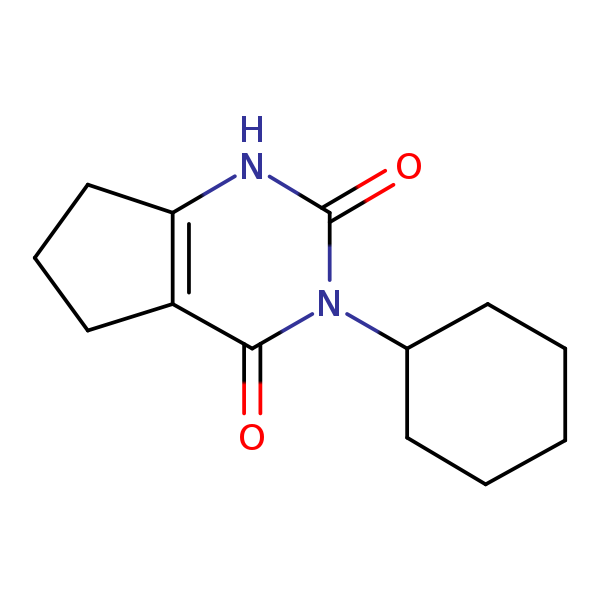 Lenacil |
Similarity: 0.63 Measured: -4.79 Predicted: -6.07 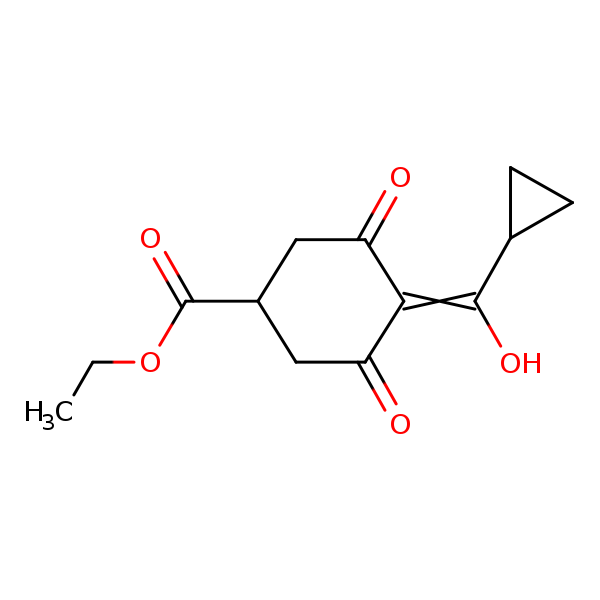 Trinexapac-ethyl |